Cancer Data Science Pulse
NCI’s ITCR Training Network Puts Cancer Research Tools and Training at Your Fingertips
Like so many areas of biomedicine today, cancer is in the midst of a data revolution. It’s now possible (and affordable) to generate large data sets in nearly all areas of cancer research and treatment. The unprecedented amount of cancer data available has led to a plethora of analysis tools, visualizations, databases, and other informatics resources, supporting nearly every aspect of cancer research and care—from basic bench research to clinical decision making. To further advance cancer research and treatment, it’s becoming increasingly necessary for investigators to have both a deep knowledge of cancer, as well as the ability to apply these specialized tools.
NCI’s Informatics Technology for Cancer Research (ITCR) program has supported the development or refinement of nearly 100 of these open-access informatics tools. Still, with data tools and technologies ever expanding, it can be a challenge to keep the larger cancer research community informed about the many resources and training opportunities available today.
The ITCR Training Network
To address this need, NCI developed an educational resource aimed at expanding training opportunities and empowering cancer researchers at all levels of experience so they may fully reap the benefits of these tools.
The goal of the ITCR Training Network (ITN) is to catalyze informatics research through training opportunities. The ITN offers training resources to enable investigators to incorporate informatics tools into their research programs and improve the usability of existing ITCR tools. Our mission is threefold and includes:
- enhancing the usability of cancer informatics tools,
- improving practices for informatics work, and
- increasing awareness of and access to informatics resources.
Courses in Development
Two courses have been launched, and others are underway. Our team is building courses based on a simple strategy; that is, keep the training user-based and as uncomplicated as possible. In other words, we’re designing the content to be used by people who are busy.
Imagine you’ve developed a new bioinformatics tool that needs to be readily understood by a broad audience. You need to be certain everyone, from entry-level graduate students to established experts in the field, can use your documentation and tutorials.
Or perhaps you’re setting up a new multidisciplinary team to perform cancer informatics research. Or maybe you’re relatively new to informatics work. Your team may include both computational and experimental students, collaborators, and employees. How do you design your studies to optimize your research goals and also ensure that each team member is properly supported?
Situations like these were the impetus behind the development of two free online courses, Documentation and Usability and Leadership for Cancer Informatics Research funded by NCI’s ITCR program. (See the ITN website for information on both courses.)
The newly launched Documentation and Usability course is targeted to informatics tool developers so they can create their own courses and learn ways to make their tools useful to the broader research community. The Leadership for Cancer Informatics Research course is geared to researchers with little or no computational experience who are building cross-disciplinary teams.
Future courses are in the works and include a broad array of topics, as listed below and described in more detail on the ITN website.
- Computing and Data Management—examines the key principles of data management from an economic, privacy, security, usability, and discoverability perspective.
- Reproducibility in Cancer Informatics and Advanced Reproducibility in Cancer Informatics—offers conceptual and hands-on training to help researchers so their findings can be easily reproduced using well-documented workflows.
- Cancer Genome Informatics—serves as an introductory course for researchers who may not be familiar with today’s informatics tools and methods, including tips for performing analyses to obtain the most rigorous results possible.
- Cancer Imaging Informatics—offers guidance for applying fully reproducible pipelines and analyses to medical and pathological images.
- Clinical Imaging Informatics—provides information on organizing and tracking clinical data, including how to extract data from clinical documents and how to summarize data across documents to save valuable time and to decrease the chance for errors when transcribing data from one format to another.
- Cancer Informatics Data Visualization—offers important principles for visualizing data. It includes a look at specific tools for visualizing omics data, as well as ideas for displaying findings from imaging and clinical information.
- Machine Learning for Cancer Informatics—examines the rapidly expanding machine learning algorithms in use today for predicting and interpreting data to advance cancer research.
- Dissemination and Engagement—provides an in-depth look at approaches for disseminating cancer informatics software, including GitHub, social media, workflow tools, and workflow publications.
Course Platforms
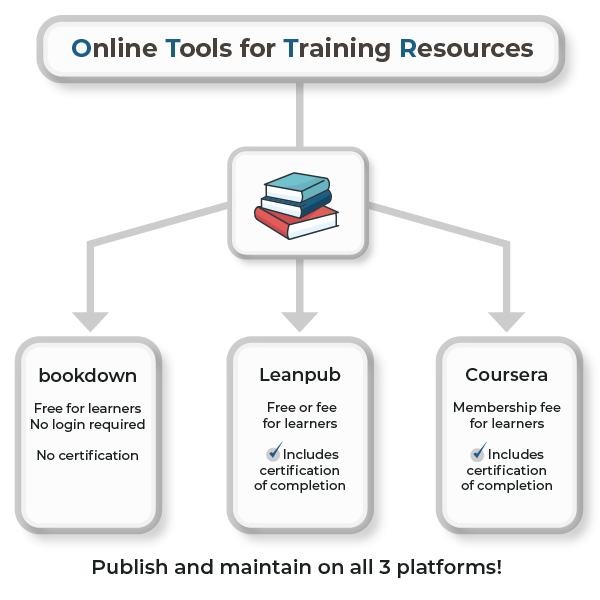
In addition to offering a wide range of training, we wanted to ensure the courses are easy to find. All the courses are published across several online educational platforms, including bookdown, Leanpub, and Coursera. Together these three platforms reach millions of learners, helping us engage users where they “live” and making it easy for them to access content.
Users log onto the site of their choice, select the course, and complete the modules. Courses are split across multiple 1-hour sections, and users can stop and restart a course at their convenience, with no time limit for completion. A full course typically takes 8–10 hours to complete. On bookdown, the course content is free (without offering a certificate). Leanpub has a pay-what-you-want option (including free) for users to access content, take quizzes, and obtain a certificate. Coursera subscribers can access the content directly within the platform and obtain a certificate after passing quizzes.
Create Your Own Course
In addition to the courses noted above, we’re encouraging members of the cancer research community to consider developing training within the Online Tools for Training Resources (OTTR) to increase accessibility to their tools.
The OTTR template offers an easy-to-create resource for developing new courses. All development is done within GitHub, an open-source environment, to allow for easy collaboration across teams. The templates help ensure courses are:
- open source.
- free to users.
- modular and mixable so content can be repackaged to address future needs.
- easy to maintain so courses can be updated to keep pace with the quickly evolving world of informatics.
Our get-started guide helps course writers navigate the powerful, but complicated, tools of GitHub and the multiple publishing platforms.
Events and Discussion
We plan to deliver course content through virtual classes and, eventually, in person when it is safe to do so. These workshops will be 2–3 hours in length and include a keynote talk from a subject matter expert, along with course content, and time for in-depth questions and demonstrations. Plans also are underway to include “train the trainer” sessions to help educators (especially at historically black colleges and universities) use the courses in their own curriculum—both to promote greater awareness about cancer informatics tools and resources and to encourage underrepresented groups to take part in cancer research. We also plan to engage community members in citizen science events to promote greater interaction between the ITCR community and the general public about cancer research.
Starting in early 2022, we’ll be offering short (about 2-hour) workshops through the Documentation and Usability course. We’ll discuss basic concepts in creating documentation and tutorials to maximize the usability of informatics tools. We’ll also introduce attendees to resources and templates to help them build or enhance their documentation, tutorials, and user feedback mechanisms. Check the ITN events page for details and to sign up for more information.
Get Involved
We encourage all members of the biomedical, bioinformatics, and IT communities to get involved in this unique one-stop resource for cancer informatics tools. Community resources like these depend on active engagement. For details on how to get involved, go to the contribute page on the ITN training website.
In addition, we welcome your ideas on the course content and presentation. You can find source material for each course in the “Future Course Materials” section of the ITCR Training Network website. You also can comment on the courses within the course source material or join our BaseCamp.
Our hope is that everyone involved in cancer research—educators, researchers, informaticists, programmers, and more—benefit from these resources, enabling them to better meet the needs of their intended audiences. Advocates for bioinformatics can help in spreading news about these tools and in refining existing resources to further advance the cancer research field.
Leave a Reply
Categories
- Data Sharing (66)
- Informatics Tools (42)
- Training (40)
- Precision Medicine (36)
- Data Standards (36)
- Genomics (36)
- Data Commons (34)
- Data Sets (27)
- Machine Learning (25)
- Artificial Intelligence (25)
- Seminar Series (22)
- Leadership Updates (14)
- Imaging (13)
- Policy (10)
- High-Performance Computing (HPC) (9)
- Jobs & Fellowships (7)
- Semantics (6)
- Funding (6)
- Proteomics (5)
- Information Technology (4)
- Awards & Recognition (3)
- Publications (2)
- Request for Information (2)
- Childhood Cancer Data Initiative (1)


Isac on June 21, 2022 at 06:47 a.m.